In the original Star Trek television episode, entitled Space Seed we are introduced to Khan Noonien Singh, a product of eugenic breeding programs on Earth. Khan and his crew are discovered by the crew of the Enterprise in deep space on a ship named the SS Botany Bay. Rescuing Khan has consequences as he and his exiled crew attempt to commandeer Enterprise to continue their original quest in building an empire led by genetically altered super beings, products of late 20th century selective breeding.

We did not fight a eugenics war in the 1990s as described in Space Seed. But we laid the foundation for creating Khans and people like him in this century. Far fetched you say? Not so much when you consider two major scientific and technological breakthroughs, products of the latter part of the 20th and the first decade of this century – the mapping of the human genome and in vitro fertilization (IVF).
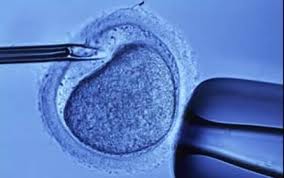
In 1978, Louise Joy Brown came into the world. Louise was no ordinary child. She was conceived in a test tube using IVF, combining a donor sperm and egg together to create an embryo and ultimately a baby. Her creators, other than the parent donors were British scientists, Dr. Patrick Steptoe, a surgeon, and Robert Edwards, a physiologist. By the time Louise turned 11 we had learned how to look inside embryos and examine gene sequences. A mere 14 years later we had a detailed map of our entire genetic makeup.
Today we have the ability to screen sperm and egg DNA and ensure that what gets combined to produce an embryo is free of inheritable traits that could cause disease in a child. The social implications of this technological capability are enormous. Parents now, if given the opportunity, have choices. They can alter the end results of natural selection using unnatural selection.
Will we in the 21st century breed Khans, a new generation of super-humans? Will we limit our ability to manipulate genes to free us of hereditary illnesses like cystic fibrosis, Tay-Sachs, sickle-cell anemia, muscular dystrophy, diabetes, blindness, deafness and heart disease? Or will we take it one step further and give parents the ability to choose eye, hair and skin colour, height and weight, sex, athletic ability and other traits having nothing to do with inherited disease?
The revolution that is unnatural selection comes with its own terminology. In this blog we will introduce you to some of it so that you can better understand what lies ahead on the road to our genetic future.
Genomics
In part 2 of this biomedicine blog I described the basic components of DNA and human genome sequencing so I would suggest you go back and read that blog before you read this next section.
We are multicellular creatures. Our cells contain nuclei (with the exception of blood cells) and each nucleus consists of paired chromosomes. We have 24 pairs in all cells except sperm and egg. Each chromosome contains linearly arranged genes. Each gene contains thousands of chemical units called base pairs, the nucleotides that combine and chain together to form DNA strands. The human genome contains 30,000 genes and 3 billion base pairs. That means on average each gene can contain 100,000 base pairs. Today we have the capability of looking at 800 base pairs at a time. So when we want to see an entire gene sequence we have to take snapshots of 800 base pair segments, overlap them and stitch the entire image together to make sure we haven’t made an error.
When we talk about an entire gene sequence we are referring to the order of the chemical unit base pairs (adenine, thymine, cytosine and guanine) in a strand of DNA. Sequencing involves breaking up a DNA strand into fragments by exposing it to free-floating nucleotides that have been assigned flourescent tags. The base pairs reassemble incorporating the flourescent tagged base units which can then be read by a computer.
In sequencing the 3 billion base pairs that make up our genetic map and comparing these maps to the sequences of people with diseases we can discover where the DNA strands have flaws. In some cases a flaw may be a base pair substitution. In other cases it could a deleted or additional base pair. In many diseases it can be a deletion of thousands of base pairs.
In developing a comprehensive database of human genome sequences from healthy and unhealthy people we have the means to create DNA screening and diagnostic testing tools.
Pre-implantation Genetic Diagnosis (PGD)
Today parents undergoing IVF can select a child’s sex and other desirable physical traits by having a doctor do PGD testing to pinpoint abnormalities in embryos, and implanting only those that are deemed healthy. PGD was first developed by Dr. Yury Verlinksy, an American, to detect potential birth defects. Today it is the standard for early prenatal diagnosis of abnormalities.
For older couples planning a pregnancy PGD testing can identify Down Syndrome, Turner Syndrome, Trisomy 18, Trisomy 13, conditions that normally do not run in a family’s medical history. PGD testing reveals specific genetic conditions like Tay Sachs, muscular dystrophy and cystic fibrosis, diseases that do run in families. In some cases an embryo can be matched to a transplant recipient such as a brother or sister suffering from leukemia by using donor stem cells and cord blood from the newborn child’s umbilicus.
Today PGD can be used to test a single gene. If you can test to eliminate any undesirable disease traits, you can also test for desired cosmetic or intellect traits and ensure that the embryos that get implanted through IVF contain the “right” genes. The implications for unnatural selection are enormous.
Preconception Screening
Why stop at just screening embryos for IVF when you can map the human genome of parents before conception? That is what preconception screening and genetic carrier screening is all about. Genetic disorders have a 25% chance of being passed to a future child when both parents carry deleted, additional or other base pair flaws in their genomes.
What is the value of preconception screening? Let’s look at one disease, cystic fibrosis. In the United States this disease affects 30,000 children and adults annually. But there are more than 8 million Americans who are asymptomatic carriers of the genes that lead to this disease. Screening could eliminate a great deal of the pain and cost associated with this inherited condition.
That’s why companies like Existence Genetics and Ambry Genetics offer genetic testing. In the case of the former, parents can take a saliva-based genetic test that screens for 1,217 rare diseases, conditions and traits. Ambry’s approach includes testing for a range of childhood diseases including a 76 disease panel test as well as individual panel tests for cystic fibrosis, Marfan Syndrome, Pancreatitis, Paranganglioma, and intellectual disabilities.
Human Germline Genetic Modification (HGGM)
This is exactly what it sounds like, changing genes in the human genome to pass along new characteristics to offspring or to correct a defect in a living person. Currently most HGGM studies are confined to laboratory mice.
HGGM is seen as a tool to be used in conjunction with IVF. An embryo can be repaired before implant, fixing a genetic mutation that could lead to disease. HGGM can also not jut fix but enhance an embryo to add athletic skills or intelligence traits. It is this enhancement capability that represents an ethical issue for many medical practitioners.
Where HGGM has application is in the treatment of defective-gene-caused diseases in living humans. The mechanism for altering the associated gene responsible for a disease involves creating a healthy gene and transferring it to the affected cells. Copies of the healthy gene are placed in a transfer agent such as a virus. The virus is inserted into affected cells where it combines with the existing DNA. The revised genome upon cell replication then is copied over and over replacing defective cells and eradicating the disease.
Of course if you extend the full capability of HGGM you can see an entire adult human genome from a somatic cell being transferred from a living person to an embryo resulting in a clone. But we’ll look at that in a separate blog.
Expect to see HGGM for IVF and disease treatment coming to a hospital or medical clinic near you in the very near future.
Pharmacogenomics
Pharmacogenomics is the study of genes and drug interactions. It looks at inherited traits in the genome and drug metabolism and response. The field combines pharmacology, biochemistry and genomics.
Drug companies are focused on developing tailor-made pharmaceuticals targeted to a specific individual’s genome. They see this as an important step in reducing adverse drug reactions in patients responsible for an estimated 100,000 deaths and 2 million hospitalizations annually in the United States alone.
Today some chemotherapy drugs such as thiopurines are not used on patients that test for an inability to produce a metabolic protein. Similar kinds of screening helps identify patients who are unable to use 3o different classes of drugs because their livers lack metabolic enzymes to breakdown the medications.
Through pharmacogenomics standard trial-and-error methods of matching patients with the right drugs will be eliminated. A physician will analyze a patient’s genome and prescribe the right drug, speeding recovery and eliminating adverse reactions.
Physicians will be able to review a patient’s genome and advise them about susceptibility to a genetic disease and include careful monitoring, diet, and lifestyle advice to lessen the impact before the disease expresses itself. Doctors will be able to provide preventive treatments to head off a genetic disease before it happens using HGGM, and vaccines designed specifically to activate the patient’s immune system.















Life Technologies have developed a benchtop DNA sequencer that will allow clinics to offer patients a genome map for $1,000. This brings DNA sequencing into the realm of everyday usage for diagnosis. See the attached press release link to learn more. http://www.lifetechnologies.com/us/en/home/about-us/news-gallery/press-releases/2012/life-techologies-itroduces-the-bechtop-io-proto.html